Abstract
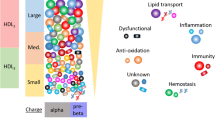
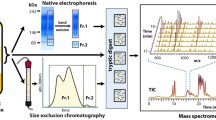
While low concentrations of high-density lipoprotein-cholesterol (HDL-C) are widely accepted as an independent cardiovascular risk factor, HDL-C-rising therapies largely failed, suggesting the importance of both HDL functions and individual subspecies. Indeed HDL particles are highly heterogeneous, with small, dense pre-beta-HDLs being considered highly biologically active but remaining poorly studied, largely reflecting difficulties for their purification. We developed an original experimental approach allowing the isolation of sufficient amounts of human pre-beta-HDLs and revealing the specificity of their proteomic and lipidomic profiles and biological activities. Pre-beta-HDLs were enriched in highly poly-unsaturated species of phosphatidic acid and phosphatidylserine, and in an unexpectedly high number of proteins implicated in the inflammatory response, including serum paraoxonase/arylesterase-1, vitronectin and clusterin, as well as in complement regulation and immunity, including haptoglobin-related protein, complement proteins and those of the immunoglobulin class. Interestingly, amongst proteins associated with lipid metabolism, phospholipid transfer protein, cholesteryl ester transfer protein and lecithin:cholesterol acyltransferase were strongly enriched in, or restricted to, pre-beta-HDL. Furthermore, pre-beta-HDL potently mediated cellular cholesterol efflux and displayed strong anti-inflammatory activities. A correlational network analysis between lipidome, proteome and biological activities highlighted 15 individual lipid and protein components of pre-beta-HDL relevant to cardiovascular disease, which may constitute novel diagnostic targets in a pathological context of altered lipoprotein metabolism.








Similar content being viewed by others
Availability of data and material
Lipidomics and proteomics data are under submissions to public databases and will be available at the date of publication.
Code availability
Not applicable.
References
Alwaili K, Bailey D, Awan Z, Bailey SD, Ruel I, Hafiane A, Krimbou L, Laboissiere S, Genest J (2012) The HDL proteome in acute coronary syndromes shifts to an inflammatory profile. Biochim Biophys Acta 1821:405–415. https://doi.org/10.1016/j.bbalip.2011.07.013
Asztalos BF, Horvath KV, Schaefer EJ (2018) High-density lipoprotein particles, cell-cholesterol efflux, and coronary heart disease risk. Arterioscler Thromb Vasc Biol 38:2007–2015. https://doi.org/10.1161/ATVBAHA.118.311117
Barrans A, Jaspard B, Barbaras R, Chap H, Perret B, Collet X (1996) Pre-beta HDL: structure and metabolism. Biochim Biophys Acta 1300:73–85
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (Methodol) 57:289–300
Camont L, Lhomme M, Rached F, Le Goff W, Nègre-Salvayre A, Salvayre R, Calzada C, Lagarde M, Chapman MJ, Kontush A (2013) Small, dense high-density lipoprotein-3 particles are enriched in negatively charged phospholipids: relevance to cellular cholesterol efflux, antioxidative, antithrombotic, anti-inflammatory, and antiapoptotic functionalities. Arterioscler Thromb Vasc Biol 33:2715–2723. https://doi.org/10.1161/ATVBAHA.113.301468
Castelli WP, Anderson K, Wilson PWF, Levy D (1992) Lipids and risk of coronary heart disease The Framingham Study. Ann Epidemiol 2:23–28. https://doi.org/10.1016/1047-2797(92)90033-M
Castro GR, Fielding CJ (1988) Early incorporation of cell-derived cholesterol into pre-beta-migrating high-density lipoprotein. Biochemistry 27:25–29
Chapman MJ, Goldstein S, Lagrange D, Laplaud PM (1981) A density gradient ultracentrifugal procedure for the isolation of the major lipoprotein classes from human serum. J Lipid Res 22:339–358
daCosta CJB, Wagg ID, McKay ME, Baenziger JE (2004) Phosphatidic acid and phosphatidylserine have distinct structural and functional interactions with the nicotinic acetylcholine receptor. J Biol Chem 279:14967–14974. https://doi.org/10.1074/jbc.M310037200
Darabi M, Guillas-Baudouin I, Le Goff W, Chapman MJ, Kontush A (2016) Therapeutic applications of reconstituted HDL: When structure meets function. Pharmacol Ther 157:28–42. https://doi.org/10.1016/j.pharmthera.2015.10.010
Darwesh AM, Sosnowski DK, Lee TY, Keshavarz-Bahaghighat H, Seubert JM (2019) Insights into the cardioprotective properties of n-3 PUFAs against ischemic heart disease via modulation of the innate immune system. Chem Biol Interact 308:20–44. https://doi.org/10.1016/j.cbi.2019.04.037
Davidson WS, Shah AS, Sexmith H, Gordon SM (2022) The HDL Proteome Watch: compilation of studies leads to new insights on HDL function. Biochim Biophys Acta Mol Cell Biol Lipids 1867:159072. https://doi.org/10.1016/j.bbalip.2021.159072
Davidson WS, Silva RAGD, Chantepie S, Lagor WR, Chapman MJ, Kontush A (2009) Proteomic analysis of defined HDL subpopulations reveals particle-specific protein clusters: relevance to antioxidative function. Arterioscler Thromb Vasc Biol 29:870–876. https://doi.org/10.1161/ATVBAHA.109.186031
Davidson WS, Sparks DL, Lund-Katz S, Phillips MC (1994) The molecular basis for the difference in charge between pre-beta- and alpha-migrating high density lipoproteins. J Biol Chem 269:8959–8965
Didichenko SA, Navdaev AV, Cukier AMO, Gille A, Schuetz P, Spycher MO, Thérond P, Chapman MJ, Kontush A, Wright SD (2016) Enhanced HDL functionality in small HDL species produced upon remodeling of HDL by reconstituted HDL, CSL112: effects on cholesterol efflux, anti-inflammatory and antioxidative activity. Circ Res 119:751–763. https://doi.org/10.1161/CIRCRESAHA.116.308685
Du X-M, Kim M-J, Hou L, Le Goff W, Chapman MJ, Van Eck M, Curtiss LK, Burnett JR, Cartland SP, Quinn CM, Kockx M, Kontush A, Rye K-A, Kritharides L, Jessup W (2015) HDL particle size is a critical determinant of ABCA1-mediated macrophage cellular cholesterol export. Circ Res 116:1133–1142. https://doi.org/10.1161/CIRCRESAHA.116.305485
Ejsing CS, Duchoslav E, Sampaio J, Simons K, Bonner R, Thiele C, Ekroos K, Shevchenko A (2006) Automated identification and quantification of glycerophospholipid molecular species by multiple precursor ion scanning. Anal Chem 78:6202–6214. https://doi.org/10.1021/ac060545x
Folch J, Lees M, Sloane Stanley GH (1957) A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem 226:497–509
Gao J, Tarcea VG, Karnovsky A, Mirel BR, Weymouth TE, Beecher CW, Cavalcoli JD, Athey BD, Omenn GS, Burant CF, Jagadish HV (2010) Metscape: a Cytoscape plug-in for visualizing and interpreting metabolomic data in the context of human metabolic networks. Bioinformatics 26:971–973. https://doi.org/10.1093/bioinformatics/btq048
Gillard BK, Rosales C, Xu B, Gotto AM, Pownall HJ (2018) Rethinking reverse cholesterol transport and dysfunctional high-density lipoproteins. J Clin Lipidol. https://doi.org/10.1016/j.jacl.2018.04.001
Harayama T, Riezman H (2018) Understanding the diversity of membrane lipid composition. Nat Rev Mol Cell Biol 19:281–296. https://doi.org/10.1038/nrm.2017.138
Hirayama S, Miida T, Miyazaki O, Aizawa Y (2007) Pre beta1-HDL concentration is a predictor of carotid atherosclerosis in type 2 diabetic patients. Diabetes Care 30:1289–1291. https://doi.org/10.2337/dc06-1948
Jorge I, Burillo E, Mesa R, Baila-Rueda L, Moreno M, Trevisan-Herraz M, Silla-Castro JC, Camafeita E, Ortega-Muñoz M, Bonzon-Kulichenko E, Calvo I, Cenarro A, Civeira F, Vázquez J (2014) The human HDL proteome displays high inter-individual variability and is altered dynamically in response to angioplasty-induced atheroma plaque rupture. J Proteom 106:61–73. https://doi.org/10.1016/j.jprot.2014.04.010
Kooijman EE, Chupin V, de Kruijff B, Burger KNJ (2003) Modulation of membrane curvature by phosphatidic acid and lysophosphatidic acid. Traffic 4:162–174
Kulig W, Korolainen H, Zatorska M, Kwolek U, Wydro P, Kepczynski M, Róg T (2019) Complex behavior of phosphatidylcholine-phosphatidic acid bilayers and monolayers: effect of acyl chain unsaturation. Langmuir 35:5944–5956. https://doi.org/10.1021/acs.langmuir.9b00381
Larrede S, Quinn CM, Jessup W, Frisdal E, Olivier M, Hsieh V, Kim M-J, Eck MV, Couvert P, Carrie A, Giral P, Chapman MJ, Guerin M, Goff WL (2009) Stimulation of cholesterol efflux by LXR agonists in cholesterol-loaded human macrophages is ABCA1-dependent but ABCG1-independent. Arterioscler Thromb Vasc Biol 29:1930–1936. https://doi.org/10.1161/ATVBAHA.109.194548
Lê Cao K-A, Martin PGP, Robert-Granié C, Besse P (2009) Sparse canonical methods for biological data integration: application to a cross-platform study. BMC Bioinform 10:34. https://doi.org/10.1186/1471-2105-10-34
Lemmon MA (2008) Membrane recognition by phospholipid-binding domains. Nat Rev Mol Cell Biol 9:99
Li L, Shi X, Guo X, Li H, Xu C (2014) Ionic protein-lipid interaction at the plasma membrane: what can the charge do? Trends Biochem Sci 39:130–140. https://doi.org/10.1016/j.tibs.2014.01.002
Lie J, de Crom R, Jauhiainen M, van Gent T, van Haperen R, Scheek L, Jansen H, Ehnholm C, van Tol A (2001) Evaluation of phospholipid transfer protein and cholesteryl ester transfer protein as contributors to the generation of pre beta-high-density lipoproteins. Biochem J 360:379–385
de la Llera-Moya M, Drazul-Schrader D, Asztalos BF, Cuchel M, Rader DJ, Rothblat GH (2010) The ability to promote efflux via ABCA1 determines the capacity of serum specimens with similar high-density lipoprotein cholesterol to remove cholesterol from macrophages. Arterioscler Thromb Vasc Biol 30:796–801. https://doi.org/10.1161/ATVBAHA.109.199158
Madsen CM, Varbo A, Nordestgaard BG (2017) Extreme high high-density lipoprotein cholesterol is paradoxically associated with high mortality in men and women: two prospective cohort studies. Eur Heart J 38:2478–2486. https://doi.org/10.1093/eurheartj/ehx163
Marsh D (1990) CRC-handbook of lipid bilayers. CRC Press, Boca Raton
Melchior JT, Street SE, Andraski AB, Furtado JD, Sacks FM, Shute RL, Greve EI, Swertfeger DK, Li H, Shah AS, Lu LJ, Davidson WS (2017) Apolipoprotein A-II alters the proteome of human lipoproteins and enhances cholesterol efflux from ABCA1. J Lipid Res 58:1374–1385. https://doi.org/10.1194/jlr.M075382
Metsalu T, Vilo J (2015) ClustVis: a web tool for visualizing clustering of multivariate data using principal component analysis and heatmap. Nucl Acids Res 43:W566-570. https://doi.org/10.1093/nar/gkv468
Miyazaki O, Ogihara J, Fukamachi I, Kasumi T (2014) Evidence for the presence of lipid-free monomolecular apolipoprotein A-1 in plasma. J Lipid Res 55:214–225. https://doi.org/10.1194/jlr.M041038
Mizuguchi C, Nakamura M, Kurimitsu N, Ohgita T, Nishitsuji K, Baba T, Shigenaga A, Shimanouchi T, Okuhira K, Otaka A, Saito H (2018) Effect of phosphatidylserine and cholesterol on membrane-mediated fibril formation by the N-terminal amyloidogenic fragment of apolipoprotein A-I. Sci Rep 8:5497. https://doi.org/10.1038/s41598-018-23920-3
Monthony JF, Wallace EG, Allen DM (1978) A non-barbital buffer for immunoelectrophoresis and zone electrophoresis in agarose gels. Clin Chem 24:1825–1827
Nanjee MN, Brinton EA (2000) Very small apolipoprotein A-I-containing particles from human plasma: isolation and quantification by high-performance size-exclusion chromatography. Clin Chem 46:207–223
Niisuke K, Kuklenyik Z, Horvath KV, Gardner MS, Toth CA, Asztalos BF (2020) Composition-function analysis of HDL subpopulations: influence of lipid composition on particle functionality. J Lipid Res. https://doi.org/10.1194/jlr.RA119000258
Pattnaik NM, Zilversmit DB (1979) Interaction of cholesteryl ester exchange protein with human plasma lipoproteins and phospholipid vesicles. J Biol Chem 254:2782–2786
Pownall HJ, Pao Q, Massey JB (1985) Acyl chain and headgroup specificity of human plasma lecithin:cholesterol acyltransferase. Separation of matrix and molecular specificities. J Biol Chem 260:2146–2152
Rached FH, Chapman MJ, Kontush A (2015) HDL particle subpopulations: focus on biological function. BioFactors 41:67–77. https://doi.org/10.1002/biof.1202
Rao R, Albers JJ, Wolfbauer G, Pownall HJ (1997) Molecular and macromolecular specificity of human plasma phospholipid transfer protein. Biochemistry 36:3645–3653. https://doi.org/10.1021/bi962776b
Rohart F, Gautier B, Singh A, Lê Cao K-A (2017) mixOmics: an R package for ’omics feature selection and multiple data integration. PLoS Comput Biol 13:e1005752. https://doi.org/10.1371/journal.pcbi.1005752
Rye K-A, Barter PJ (2014) Regulation of high-density lipoprotein metabolism. Circ Res 114:143–156. https://doi.org/10.1161/CIRCRESAHA.114.300632
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J (2003) TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34:374–378. https://doi.org/10.2144/03342mt01
Serfaty-Lacrosniere C, Civeira F, Lanzberg A, Isaia P, Berg J, Janus ED, Smith MP, Pritchard PH, Frohlich J, Lees RS (1994) Homozygous Tangier disease and cardiovascular disease. Atherosclerosis 107:85–98. https://doi.org/10.1016/0021-9150(94)90144-9
Shah AS, Tan L, Long JL, Davidson WS (2013) Proteomic diversity of high density lipoproteins: our emerging understanding of its importance in lipid transport and beyond. J Lipid Res 54:2575–2585. https://doi.org/10.1194/jlr.R035725
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Singh A, Shannon CP, Gautier B, Rohart F, Vacher M, Tebbutt SJ, Lê Cao K-A (2019) DIABLO: an integrative approach for identifying key molecular drivers from multi-omics assays. Bioinformatics 35:3055–3062. https://doi.org/10.1093/bioinformatics/bty1054
Stace CL, Ktistakis NT (2006) Phosphatidic acid- and phosphatidylserine-binding proteins. Biochimica et Biophysica Acta (BBA) Mol Cell Biol Lipids 1761:913–926. https://doi.org/10.1016/j.bbalip.2006.03.006
Suzuki M, Pritchard DK, Becker L, Hoofnagle AN, Tanimura N, Bammler TK, Beyer RP, Bumgarner R, Vaisar T, de Beer MC, de Beer FC, Miyake K, Oram JF, Heinecke JW (2010) High-density lipoprotein suppresses the type I interferon response, a family of potent antiviral immunoregulators, in macrophages challenged with lipopolysaccharide. Circulation 122:1919–1927. https://doi.org/10.1161/CIRCULATIONAHA.110.961193
Swertfeger DK, Li H, Rebholz S, Zhu X, Shah AS, Davidson WS, Lu LJ (2017) Mapping atheroprotective functions and related proteins/lipoproteins in size fractionated human plasma. Mol Cell Proteomics 16:680–693. https://doi.org/10.1074/mcp.M116.066290
Tall AR, Forester LR, Bongiovanni GL (1983) Facilitation of phosphatidylcholine transfer into high density lipoproteins by an apolipoprotein in the density 1.20-1.26 g/ml fraction of plasma. J Lipid Res 24:277–289
Vaisar T, Pennathur S, Green PS, Gharib SA, Hoofnagle AN, Cheung MC, Byun J, Vuletic S, Kassim S, Singh P, Chea H, Knopp RH, Brunzell J, Geary R, Chait A, Zhao X-Q, Elkon K, Marcovina S, Ridker P, Oram JF, Heinecke JW (2007) Shotgun proteomics implicates protease inhibition and complement activation in the antiinflammatory properties of HDL. J Clin Investig 117:746–756. https://doi.org/10.1172/JCI26206
Weckerle A, Snipes JA, Cheng D, Gebre AK, Reisz JA, Murea M, Shelness GS, Hawkins GA, Furdui CM, Freedman BI, Parks JS, Ma L (2016) Characterization of circulating APOL1 protein complexes in African Americans. J Lipid Res 57:120–130. https://doi.org/10.1194/jlr.M063453
Yoon M-S, Rosenberger CL, Wu C, Truong N, Sweedler JV, Chen J (2015) Rapid mitogenic regulation of the mTORC1 inhibitor, DEPTOR, by phosphatidic acid. Mol Cell 58:549–556. https://doi.org/10.1016/j.molcel.2015.03.028
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: these studies were supported by National Institute for Health and Medical Research (INSERM; Paris, France) and Sorbonne University (Paris, France). We gratefully acknowledge further support from ICAN (Paris, France) and China Scholarship Council (CSC, China).
Author information
Authors and Affiliations
Contributions
IG, AK, and WLG designed the experiments; IG, MD, FM and MD performed the experiments; IG and SL performed the LPLC experiments; ML performed the lipidomic and CP and LM the proteomic analysis; PC, PG, PEK and MA performed the clinical assessments, and Tangier patient recruitment; MP performed the statistical and bioinformatics analysis; IG, AK, and WLG analysed the data; IG, PL, MG, AK, and WLG wrote the manuscript. All the authors gave critical comments on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: AK reports grants from CSL and Pfizer, personal fees from American Heart Association, National Lipid Association, Sanofi and European Society of Cardiology, outside the submitted work.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guillas, I., Lhomme, M., Pionneau, C. et al. Identification of the specific molecular and functional signatures of pre-beta-HDL: relevance to cardiovascular disease. Basic Res Cardiol 118, 33 (2023). https://doi.org/10.1007/s00395-023-01004-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00395-023-01004-2