Abstract
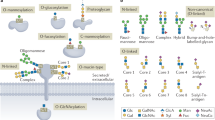
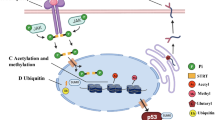
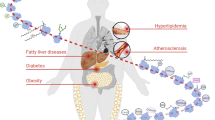
Protein glycosylation, which involves the attachment of carbohydrates to proteins, is one of the most abundant protein co-translational and post-translational modifications. Advances in technology have substantially increased our knowledge of the biosynthetic pathways involved in protein glycosylation, as well as how changes in glycosylation can affect cell function. In addition, our understanding of the role of protein glycosylation in disease processes is growing, particularly in the context of immune system function, infectious diseases, neurodegeneration and cancer. Several decades ago, cell surface glycoproteins were found to have an important role in regulating ion transport across the cardiac sarcolemma. However, with very few exceptions, our understanding of how changes in protein glycosylation influence cardiovascular (patho)physiology remains remarkably limited. Therefore, in this Review, we aim to provide an overview of N-linked and O-linked protein glycosylation, including intracellular O-linked N-acetylglucosamine protein modification. We discuss our current understanding of how all forms of protein glycosylation contribute to normal cardiovascular function and their roles in cardiovascular disease. Finally, we highlight potential gaps in our knowledge about the effects of protein glycosylation on the heart and vascular system, highlighting areas for future research.
Key points
-
Protein glycosylation is one of the most abundant protein modifications, affecting secreted, extracellular and intracellular proteins.
-
In the cardiovascular system, protein glycosylation regulates diverse homeostatic functions, including contractility, metabolism and transcription, as well as extracellular and intracellular signalling pathways.
-
In the heart, regulation of protein glycosylation contributes to the pathophysiology of heart failure and cardiac hypertrophy and the adverse effects of diabetes mellitus.
-
Dysregulation of protein glycosylation in the vascular system contributes to changes in cell–cell interactions, inflammation and vascular remodelling, including atherosclerosis.
-
Opportunities for future research include the development of humanized mouse models, improved understanding of the regulation of glycogene expression, application of single-cell techniques, and novel approaches to modulate N-glycoforms in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Khoury, G. A., Baliban, R. C. & Floudas, C. A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the Swiss-Prot database. Sci. Rep. 1, 90 (2011).
Apweiler, R., Hermjakob, H. & Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta 1473, 4–8 (1999).
Hart, G. W. & Copeland, R. J. Glycomics hits the big time. Cell 143, 672–676 (2010).
Cummings, R. D. & Pierce, J. M. The challenge and promise of glycomics. Chem. Biol. 21, 1–15 (2014).
Schjoldager, K. T., Narimatsu, Y., Joshi, H. J. & Clausen, H. Global view of human protein glycosylation pathways and functions. Nat. Rev. Mol. Cell Biol. 21, 729–749 (2020).
Varki, A. Biological roles of glycans. Glycobiology 27, 3–49 (2017).
Bennett, H. S. Morphological aspects of extracellular polysaccharides. J. Hist. Cytochem. 11, 14–23 (1963).
Cook, G. M. Glycoproteins in membranes. Biol. Rev. Camb. Philos. Soc. 43, 363–391 (1968).
Spiro, R. G. Glycoproteins: structure, metabolism and biology. N. Engl. J. Med. 269, 616–621 (1963).
Gee, D. J. A glycoprotein in cardiac conducting tissue. Br. Heart J. 31, 588–590 (1969).
Langer, G. A., Frank, J. S., Nudd, L. M. & Seraydarian, K. Sialic acid: effect of removal on calcium exchangeability of cultured heart cells. Science 193, 1013–1015 (1976).
Frank, J. S., Langer, G. A., Nudd, L. M. & Seraydarian, K. The myocardial cell surface, its histochemistry, and the effect of sialic acid and calcium removal on its stucture and cellular ionic exchange. Circ. Res. 41, 702–714 (1977).
Varki, A. et al. (eds) Essentials of Glycobiology 4th edn (Cold Spring Harbor Laboratory Press, 2022).
Varki, A. & Kornfeld, S. in Essentials of Glycobiology 4th edn (Varki, A. et al. eds) 1–20 (Cold Spring Harbor Laboratory Press, 2022).
Haltiwanger, R. S. et al. in Essentials of Glycobiology 4th edn (Varki, A. et al. eds) 155–164 (Cold Spring Harbor Laboratory Press, 2022).
Minakata, S. et al. Protein C-mannosylation and C-mannosyl tryptophan in chemical biology and medicine. Molecules 26, 5258 (2021).
Haynes, P. A. Phosphoglycosylation: a new structural class of glycosylation? Glycobiology 8, 1–5 (1998).
Maynard, J. C., Burlingame, A. L. & Medzihradszky, K. F. Cysteine S-linked N-acetylglucosamine (S-GlcNAcylation), a new post-translational modification in mammals. Mol. Cell Proteom. 15, 3405–3411 (2016).
Stanley, P., Moremen, K. W., Lewis, N. E, Taniguchi, N. & Aebi, M. N-Glycans. In Essentials of Glycobiology 4th edn (eds Varki, A. et al.) 103–116 (Cold Spring Harbor Laboratory Press, 2022).
Lewis, A. L., Chen X., Schnaar, R. L. & Varki, A. Sialic acids and other nonulosonic acids. In Essentials of Glycobiology 4th edn (eds Varki, A. et al.) 185–204 (Cold Spring Harbor Laboratory Press, 2022).
Brockhausen, I., Wandall, H. H., Ten Hagen, K. G. & Stanley, P. O-GalNAc glycans. In Essentials of Glycobiology 4th edn (eds Varki, A. et al.) 117–128 (Cold Spring Harbor Laboratory Press, 2022).
Reily, C., Stewart, T. J., Renfrow, M. B. & Novak, J. Glycosylation in health and disease. Nat. Rev. Nephrol. 15, 346–366 (2019).
Merry, C. L. R., Lindahl, U, Couchman, J. & Esko, J. D. Proteoglycans and sulfated glycosaminoglycans. In Essentials of Glycobiology 4th edn (eds Varki, A. et al.) 217–232 (Cold Spring Harbor Laboratory Press, 2022).
Iozzo, R. V. & Schaefer, L. Proteoglycan form and function: a comprehensive nomenclature of proteoglycans. Matrix Biol. 42, 11–55 (2015).
Christensen, G., Herum, K. M. & Lunde, I. G. Sweet, yet underappreciated: proteoglycans and extracellular matrix remodeling in heart disease. Matrix Biol. 75-76, 286–299 (2019).
Frangogiannis, N. G. The extracellular matrix in ischemic and nonischemic heart failure. Circ. Res. 125, 117–146 (2019).
Rienks, M., Papageorgiou, A. P., Frangogiannis, N. G. & Heymans, S. Myocardial extracellular matrix: an ever-changing and diverse entity. Circ. Res. 114, 872–888 (2014).
Zimmer, B. M., Barycki, J. J. & Simpson, M. A. Mechanisms of coordinating hyaluronan and glycosaminoglycan production by nucleotide sugars. Am. J. Physiol. Cell Physiol. 322, C1201–C1213 (2022).
Caon, I. et al. Cell energy metabolism and hyaluronan synthesis. J. Histochem. Cytochem. 69, 35–47 (2021).
Torres, C. R. & Hart, G. W. Topography and polypeptide distribution of terminal N-acetylglucosamine residues on the surfaces of intact lymphocytes. Evidence for O-linked GlcNAc. J. Biol. Chem. 259, 3308–3317 (1984).
Holt, G. D. & Hart, G. W. The subcellular distribution of terminal N-acetylglucosamine moieties. Localization of a novel protein-saccharide linkage, O-linked GlcNAc. J. Biol. Chem. 261, 8049–8057 (1986).
Chatham, J. C., Zhang, J. & Wende, A. R. Role of O-linked N-acetylglucosamine protein modification in cellular (patho)physiology. Physiol. Rev. 101, 427–493 (2021).
Matsuura, A. et al. O-linked N-acetylglucosamine is present on the extracellular domain of notch receptors. J. Biol. Chem. 283, 35486–35495 (2008).
Ogawa, M. & Okajima, T. Structure and function of extracellular O-GlcNAc. Curr. Opin. Struct. Biol. 56, 72–77 (2019).
Ogawa, M., Senoo, Y., Ikeda, K., Takeuchi, H. & Okajima, T. Structural divergence in O-GlcNAc glycans displayed on epidermal growth factor-like repeats of mammalian notch1. Molecules 23, 1745 (2018).
Varshney, S. & Stanley, P. EOGT and O-GlcNAc on secreted and membrane proteins. Biochem. Soc. Trans. 45, 401–408 (2017).
Shaheen, R. et al. Mutations in EOGT confirm the genetic heterogeneity of autosomal-recessive Adams–Oliver syndrome. Am. J. Hum. Genet. 92, 598–604 (2013).
Cohen, I. et al. Autosomal recessive Adams–Oliver syndrome caused by homozygous mutation in EOGT, encoding an EGF domain-specific O-GlcNAc transferase. Eur. J. Hum. Genet. 22, 374–378 (2014).
Sawaguchi, S. et al. O-GlcNAc on NOTCH1 EGF repeats regulates ligand-induced Notch signaling and vascular development in mammals. Elife 6, e24419 (2017).
DeHaven, J. E., Robinson, K. A., Nelson, B. A. & Buse, M. G. A novel variant of glutamine: fructose-6-phosphate amidotransferase-1 (GFAT1) mRNA is selectively expressed in striated muscle. Diabetes 50, 2419–2424 (2001).
Liu, K. et al. Molecular characterization, chromosomal location, alternative splicing and polymorphism of porcine GFAT1 gene. Mol. Biol. Rep. 37, 2711–2717 (2010).
Wang, Z. V. et al. Spliced X-box binding protein 1 couples the unfolded protein response to hexosamine biosynthetic pathway. Cell 156, 1179–1192 (2014).
Nabeebaccus, A. A. et al. Cardiomyocyte protein O-GlcNAcylation is regulated by GFAT1 not GFAT2. Biochem. Biophys. Res. Commun. 583, 121–127 (2021).
Ishikita, A. et al. GFAT2 mediates cardiac hypertrophy through HBP-O-GlcNAcylation-Akt pathway. iScience 24, 103517 (2021).
Freeze, H. H. et al. Glycosylation precursors. In Essentials of Glycobiology (eds Varki, A. et al.) 53–66 (Cold Spring Harbor Laboratory Press, 2022).
Harduin-Lepers, A. The vertebrate sialylation machinery: structure-function and molecular evolution of GT-29 sialyltransferases. Glycoconj. J. 40, 473–492 (2023).
Kreppel, L. K. & Hart, G. W. Regulation of a cytosolic and nuclear O-GlcNAc transferase. Role of the tetratricopeptide repeats. J. Biol. Chem. 274, 32015–32022 (1999).
Dong, Y. Y. et al. Structures of DPAGT1 explain glycosylation disease mechanisms and advance TB antibiotic design. Cell 175, 1045–1058 (2018).
Song, W. et al. O-GlcNAcylation regulates β1,4-GlcNAc-branched N-glycan biosynthesis via the OGT/SLC35A3/GnT-IV axis. FASEB J. 36, e22149 (2022).
Vella, P. et al. Tet proteins connect the O-linked N-acetylglucosamine transferase OGT to chromatin in embryonic stem cells. Mol. Cell 49, 645–656 (2013).
Vigetti, D. et al. Role of UDP-N-acetylglucosamine (GlcNAc) and O-GlcNAcylation of hyaluronan synthase 2 in the control of chondroitin sulfate and hyaluronan synthesis. J. Biol. Chem. 287, 35544–35555 (2012).
Bennmann, D., Weidemann, W., Thate, A., Kreuzmann, D. & Horstkorte, R. Aberrant O-GlcNAcylation disrupts GNE enzyme activity in GNE myopathy. FEBS J. 283, 2285–2294 (2016).
Ghosh, S. K. et al. Disruption of O-GlcNAc cycling in C. elegans perturbs nucleotide sugar pools and complex glycans. Front. Endocrinol. 5, 197 (2014).
Marques-da-Silva, D. et al. Cardiac complications of congenital disorders of glycosylation (CDG): a systematic review of the literature. J. Inherit. Metab. Dis. 40, 657–672 (2017).
Montpetit, M. L. et al. Regulated and aberrant glycosylation modulate cardiac electrical signaling. Proc. Natl Acad. Sci. USA 106, 16517–16522 (2009).
Franzka, P. et al. Altered glycosylation in the aging heart. Front. Mol. Biosci. 8, 673044 (2021).
Watson, L. J. et al. Cardiomyocyte Ogt is essential for postnatal viability. Am. J. Physiol. Heart Circ. Physiol. 306, H142–H153 (2014).
Mu, Y. et al. O-linked β-N-acetylglucosamine transferase plays an essential role in heart development through regulating angiopoietin-1. PLoS Genet. 16, e1008730 (2020).
Dassanayaka, S. et al. Cardiomyocyte Oga haploinsufficiency increases O-GlcNAcylation but hastens ventricular dysfunction following myocardial infarction. PLoS ONE 15, e0242250 (2020).
Ha, C.-M. et al. Sustained O-GlcNAc levels leads to cardiac hypertrophy and reduced mitochondrial function without systolic contractile impairment. J. Am. Heart Assoc. 12, e029898 (2023).
Umapathi, P. et al. Excessive O-GlcNAcylation causes heart failure and sudden death. Circulation 143, 1687–1703 (2021).
Laczy, B., Marsh, S. A., Brocks, C. A., Wittmann, I. & Chatham, J. C. Inhibition of O-GlcNAcase in perfused rat hearts by NAG-thiazolines at the time of reperfusion is cardioprotective in an O-GlcNAc-dependent manner. Am. J. Physiol. Heart circ. Physiol. 299, H1715–H1727 (2010).
Ngoh, G. A., Watson, L. J., Facundo, H. T., Dillmann, W. & Jones, S. P. Non-canonical glycosyltransferase modulates post-hypoxic cardiac myocyte death and mitochondrial permeability transition. J. Mol. Cell. Cardiol. 45, 313–325 (2008).
Jones, S. P. et al. Cardioprotection by N-acetylglucosamine linkage to cellular proteins. Circulation 117, 1172–1182 (2008).
Narayanan, B. et al. Cardioprotective O-GlcNAc signaling is elevated in murine female hearts via enhanced O-GlcNAc transferase activity. J. Biol. Chem. 299, 105447 (2023).
Ufret-Vincenty, C. A. et al. Role of sodium channel deglycosylation in the genesis of cardiac arrhythmias in heart failure. J. Biol. Chem. 276, 28197–28203 (2001).
Ednie, A. R., Horton, K. K., Wu, J. & Bennett, E. S. Expression of the sialyltransferase, ST3Gal4, impacts cardiac voltage-gated sodium channel activity, refractory period and ventricular conduction. J. Mol. Cell. Cardiol. 59, 117–127 (2013).
Ednie, A. R. & Bennett, E. S. Reduced sialylation impacts ventricular repolarization by modulating specific K+ channel isoforms distinctly. J. Biol. Chem. 290, 2769–2783 (2015).
Jay, S. D. et al. Structural characterization of the dihydropyridine-sensitive calcium channel alpha 2-subunit and the associated delta peptides. J. Biol. Chem. 266, 3287–3293 (1991).
Tetreault, M. P. et al. Identification of glycosylation sites essential for surface expression of the CaVα2δ1 subunit and modulation of the cardiac CaV1.2 channel activity. J. Biol. Chem. 291, 4826–4843 (2016).
Weiss, N., Black, S. A., Bladen, C., Chen, L. & Zamponi, G. W. Surface expression and function of Cav3.2 T-type calcium channels are controlled by asparagine-linked glycosylation. Pflug. Arch. 465, 1159–1170 (2013).
Ondacova, K., Karmazinova, M., Lazniewska, J., Weiss, N. & Lacinova, L. Modulation of Cav3.2 T-type calcium channel permeability by asparagine-linked glycosylation. Channels 10, 175–184 (2016).
Liu, Y. et al. Asparagine-linked glycosylation modifies voltage-dependent gating properties of CaV3.1-T-type Ca2+ channel. J. Physiol. Sci. 69, 335–343 (2019).
Schwetz, T. A., Norring, S. A., Ednie, A. R. & Bennett, E. S. Sialic acids attached to O-glycans modulate voltage-gated potassium channel gating. J. Biol. Chem. 286, 4123–4132 (2011).
Niwa, N. & Nerbonne, J. M. Molecular determinants of cardiac transient outward potassium current (Ito) expression and regulation. J. Mol. Cell Cardiol. 48, 12–25 (2010).
Ednie, A. R., Harper, J. M. & Bennett, E. S. Sialic acids attached to N- and O-glycans within the Nav1.4 D1S5-S6 linker contribute to channel gating. Biochim. Biophys. Acta 1850, 307–317 (2015).
Chandrasekhar, K. D. et al. O-glycosylation of the cardiac IKs complex. J. Physiol. 589, 3721–3730 (2011).
Yu, P. et al. O-GlcNAcylation of cardiac Nav1.5 contributes to the development of arrhythmias in diabetic hearts. Int. J. Cardiol. 260, 74–81 (2018).
Ednie, A. R. & Bennett, E. S. Intracellular O-linked glycosylation directly regulates cardiomyocyte L-type Ca2+ channel activity and excitation-contraction coupling. Basic. Res. Cardiol. 115, 59 (2020).
Ednie, A. R., Paul-Onyia, C. D. & Bennett, E. S. Reduced O-GlcNAcylation diminishes cardiomyocyte Ca2+ dependent facilitation and frequency dependent acceleration of relaxation. J. Mol. Cell Cardiol. 180, 10–21 (2023).
Hegyi, B. et al. Hyperglycemia regulates cardiac K+ channels via O-GlcNAc-CaMKII and NOX2-ROS-PKC pathways. Basic. Res. Cardiol. 115, 71 (2020).
Okolo, C. A. et al. Direct regulation of the cardiac ryanodine receptor (RyR2) by O-GlcNAcylation. Cardiovasc. Diabetol. 22, 276 (2023).
Goth, C. K., Petaja-Repo, U. E. & Rosenkilde, M. M. G protein-coupled receptors in the sweet spot: glycosylation and other post-translational modifications. ACS Pharmacol. Transl. Sci. 3, 237–245 (2020).
Rands, E. et al. Mutational analysis of beta-adrenergic receptor glycosylation. J. Biol. Chem. 265, 10759–10764 (1990).
Hakalahti, A. E. et al. Human β1-adrenergic receptor is subject to constitutive and regulated N-terminal cleavage. J. Biol. Chem. 285, 28850–28861 (2010).
He, J., Xu, J., Castleberry, A. M., Lau, A. G. & Hall, R. A. Glycosylation of β1-adrenergic receptors regulates receptor surface expression and dimerization. Biochem. Biophys. Res. Commun. 297, 565–572 (2002).
Park, M., Reddy, G. R., Wallukat, G., Xiang, Y. K. & Steinberg, S. F. β1-adrenergic receptor O-glycosylation regulates N-terminal cleavage and signaling responses in cardiomyocytes. Sci. Rep. 7, 7890 (2017).
Goth, C. K. et al. Site-specific O-glycosylation by polypeptide N-acetylgalactosaminyltransferase 2 (GalNAc-transferase T2) co-regulates β1-adrenergic receptor N-terminal cleavage. J. Biol. Chem. 292, 4714–4726 (2017).
Cao, H. et al. O-GlcNAc transferase affects the signal transduction of β1 adrenoceptor in adult rat cardiomyocytes by increasing the O-GlcNAcylation of β1 adrenoceptor. Biochem. Biophys. Res. Commun. 528, 71–77 (2020).
Woo, C. M. et al. Mapping and quantification of over 2000 O-linked glycopeptides in activated human t cells with isotope-targeted glycoproteomics (Isotag). Mol. Cell Proteom. 17, 764–775 (2018).
Zhang, J., Simpson, P. C. & Jensen, B. C. Cardiac α1A-adrenergic receptors: emerging protective roles in cardiovascular diseases. Am. J. Physiol. Heart circ. Physiol. 320, H725–H733 (2021).
Hynes, R. O. & Naba, A. Overview of the matrisome – an inventory of extracellular matrix constituents and functions. Cold Spring Harb. Perspect. Biol. 4, a004903 (2012).
Halper, J. Basic components of connective tissues and extracellular matrix: fibronectin, fibrinogen, laminin, elastin, fibrillins, fibulins, matrilins, tenascins and thrombospondins. Adv. Exp. Med. Biol. 1348, 105–126 (2021).
Adams, J. C. & Lawler, J. The thrombospondins. Cold Spring Harb. Perspect. Biol. 3, a009712 (2011).
Forbes, T., Pauza, A. G. & Adams, J. C. In the balance: how do thrombospondins contribute to the cellular pathophysiology of cardiovascular disease? Am. J. Physiol. Cell Physiol. 321, C826–C845 (2021).
Swinnen, M. et al. Absence of thrombospondin-2 causes age-related dilated cardiomyopathy. Circulation 120, 1585–1597 (2009).
Matsumoto, K. I. & Aoki, H. The roles of tenascins in cardiovascular, inflammatory, and heritable connective tissue diseases. Front. Immunol. 11, 609752 (2020).
Frangogiannis, N. G. & Kovacic, J. C. Extracellular matrix in ischemic heart disease, part 4/4: JACC focus seminar. J. Am. Coll. Cardiol. 75, 2219–2235 (2020).
Diez, J., Gonzalez, A. & Kovacic, J. C. Myocardial interstitial fibrosis in nonischemic heart disease, part 3/4: JACC focus seminar. J. Am. Coll. Cardiol. 75, 2204–2218 (2020).
Del Monte-Nieto, G., Fischer, J. W., Gorski, D. J., Harvey, R. P. & Kovacic, J. C. Basic biology of extracellular matrix in the cardiovascular system, part 1/4: JACC focus seminar. J. Am. Coll. Cardiol. 75, 2169–2188 (2020).
Oka, T. et al. Genetic manipulation of periostin expression reveals a role in cardiac hypertrophy and ventricular remodeling. Circ. Res. 101, 313–321 (2007).
Shimazaki, M. et al. Periostin is essential for cardiac healing after acute myocardial infarction. J. Exp. Med. 205, 295–303 (2008).
Schellings, M. W. et al. Absence of SPARC results in increased cardiac rupture and dysfunction after acute myocardial infarction. J. Exp. Med. 206, 113–123 (2009).
Bradshaw, A. D. et al. Pressure overload-induced alterations in fibrillar collagen content and myocardial diastolic function: role of secreted protein acidic and rich in cysteine (SPARC) in post-synthetic procollagen processing. Circulation 119, 269–280 (2009).
Heidenreich, P. A. et al. 2022 AHA/ACC/HFSA guideline for the management of heart failure: executive summary: a report of the American College of Cardiology/American Heart Association Joint Committee on Clinical Practice Guidelines. Circulation 145, e876–e894 (2022).
Ednie, A. R., Deng, W., Yip, K. P. & Bennett, E. S. Reduced myocyte complex N-glycosylation causes dilated cardiomyopathy. FASEB J. 33, 1248–1261 (2019).
Deng, W., Ednie, A. R., Qi, J. & Bennett, E. S. Aberrant sialylation causes dilated cardiomyopathy and stress-induced heart failure. Basic. Res. Cardiol. 111, 57 (2016).
Kiarash, A. et al. Defective glycosylation of calsequestrin in heart failure. Cardiovasc. Res. 63, 264–272 (2004).
Jacob, S. et al. Altered calsequestrin glycan processing is common to diverse models of canine heart failure. Mol. Cell Biochem. 377, 11–21 (2013).
Lapidos, K. A., Kakkar, R. & McNally, E. M. The dystrophin glycoprotein complex: signaling strength and integrity for the sarcolemma. Circ. Res. 94, 1023–1031 (2004).
Mestroni, L. et al. Familial dilated cardiomyopathy: evidence for genetic and phenotypic heterogeneity. Heart Muscle Disease Study Group. J. Am. Coll. Cardiol. 34, 181–190 (1999).
Townsend, D. Finding the sweet spot: assembly and glycosylation of the dystrophin-associated glycoprotein complex. Anat. Rec. 297, 1694–1705 (2014).
Endo, T. Glycobiology of α-dystroglycan and muscular dystrophy. J. Biochem. 157, 1–12 (2015).
Valera, I. C. et al. Essential roles of the dystrophin-glycoprotein complex in different cardiac pathologies. Adv. Med. Sci. 66, 52–71 (2021).
Ujihara, Y. et al. Elimination of fukutin reveals cellular and molecular pathomechanisms in muscular dystrophy-associated heart failure. Nat. Commun. 10, 5754 (2019).
Michele, D. E., Kabaeva, Z., Davis, S. L., Weiss, R. M. & Campbell, K. P. Dystroglycan matrix receptor function in cardiac myocytes is important for limiting activity-induced myocardial damage. Circ. Res. 105, 984–993 (2009).
Barresi, R. et al. Disruption of heart sarcoglycan complex and severe cardiomyopathy caused by β sarcoglycan mutations. J. Med. Genet. 37, 102–107 (2000).
Fayssoil, A., Nardi, O., Annane, D. & Orlikowski, D. Left ventricular function in alpha-sarcoglycanopathy and gamma-sarcoglycanopathy. Acta Neurol. Belg. 114, 257–259 (2014).
Calvo, F. et al. Evaluation of heart involvement in gamma-sarcoglycanopathy (LGMD2C-). A study of ten patients. Neuromuscul. Disord. 10, 560–566 (2000).
Alonso-Perez, J. et al. Clinical and genetic spectrum of a large cohort of patients with δ-sarcoglycan muscular dystrophy. Brain 145, 596–606 (2022).
Schade van Westrum, S. M. et al. Cardiac involvement in Dutch patients with sarcoglycanopathy: a cross-sectional cohort and follow-up study. Muscle Nerve 50, 909–913 (2014).
Sveen, M. L., Thune, J. J., Kober, L. & Vissing, J. Cardiac involvement in patients with limb-girdle muscular dystrophy type 2 and Becker muscular dystrophy. Arch. Neurol. 65, 1196–1201 (2008).
Durbeej, M. et al. Disruption of the β-sarcoglycan gene reveals pathogenetic complexity of limb-girdle muscular dystrophy type 2E. Mol. Cell 5, 141–151 (2000).
Coral-Vazquez, R. et al. Disruption of the sarcoglycan-sarcospan complex in vascular smooth muscle: a novel mechanism for cardiomyopathy and muscular dystrophy. Cell 98, 465–474 (1999).
Hack, A. A. et al. Gamma-sarcoglycan deficiency leads to muscle membrane defects and apoptosis independent of dystrophin. J. Cell Biol. 142, 1279–1287 (1998).
Duclos, F. et al. Progressive muscular dystrophy in α-sarcoglycan-deficient mice. J. Cell Biol. 142, 1461–1471 (1998).
Young, M. E. et al. Proposed regulation of gene expression by glucose in rodent heart. Gene Regul. Syst. Bio 1, 251–262 (2007).
Yang, S. et al. Integrated glycoprotein immobilization method for glycopeptide and glycan analysis of cardiac hypertrophy. Anal. Chem. 87, 9671–9678 (2015).
Rong, J. et al. Glycan imaging in intact rat hearts and glycoproteomic analysis reveal the upregulation of sialylation during cardiac hypertrophy. J. Am. Chem. Soc. 136, 17468–17476 (2014).
Nagai-Okatani, C. & Minamino, N. Aberrant glycosylation in the left ventricle and plasma of rats with cardiac hypertrophy and heart failure. PLoS ONE 11, e0150210 (2016).
Lunde, I. G. et al. Cardiac O-GlcNAc signaling is increased in hypertrophy and heart failure. Physiol. Genomics 44, 162–172 (2012).
Agrawal, P. et al. Mapping posttranscriptional regulation of the human glycome uncovers microRNA defining the glycocode. Proc. Natl Acad. Sci. USA 111, 4338–4343 (2014).
Wehbe, N. et al. MicroRNAs in cardiac hypertrophy. Int. J. Mol. Sci. 20, 4714 (2019).
Indellicato, R. & Trinchera, M. Epigenetic regulation of glycosylation in cancer and other diseases. Int. J. Mol. Sci. 22, 2980 (2021).
Qin, J., Guo, N., Tong, J. & Wang, Z. Function of histone methylation and acetylation modifiers in cardiac hypertrophy. J. Mol. Cell. Cardiol. 159, 120–129 (2021).
Mailleux, F., Gélinas, R., Beauloye, C., Horman, S. & Bertrand, L. O-GlcNAcylation, enemy or ally during cardiac hypertrophy development? BBA Mol. Basis Dis. 1862, 2232–2243 (2016).
Facundo, H. T. et al. O-GlcNAc signaling is essential for NFAT-mediated transcriptional reprogramming during cardiomyocyte hypertrophy. Am. J. Physiol. Heart Circ. Physiol. 302, H2122–H2130 (2012).
Gelinas, R. et al. AMPK activation counteracts cardiac hypertrophy by reducing O-GlcNAcylation. Nat. Commun. 9, 374 (2018).
Tran, D. H. et al. Chronic activation of hexosamine biosynthesis in the heart triggers pathological cardiac remodeling. Nat. Commun. 11, 1771 (2020).
Zhu, W. Z., El-Nachef, D., Yang, X., Ledee, D. & Olson, A. K. O-GlcNAc transferase promotes compensated cardiac function and protein kinase A O-GlcNAcylation during early and established pathological hypertrophy from pressure overload. J. Am. Heart Assoc. 8, e011260 (2019).
Prakoso, D. et al. Fine-tuning the cardiac O-GlcNAcylation regulatory enzymes governs the functional and structural phenotype of the diabetic heart. Cardiovasc. Res. 118, 212–225 (2022).
Zhu, W. Z. et al. First comprehensive identification of cardiac proteins with putative increased O-GlcNAc levels during pressure overload hypertrophy. PLoS ONE 17, e0276285 (2022).
Ritchie, R. H. & Abel, E. D. Basic mechanisms of diabetic heart disease. Circ. Res. 126, 1501–1525 (2020).
Zhao, Q., Jia, T. Z., Cao, Q. C., Tian, F. & Ying, W. T. A crude 1-DNJ extract from home made Bombyx batryticatus inhibits diabetic cardiomyopathy-associated fibrosis in db/db mice and reduces protein N-glycosylation levels. Int. J. Mol. Sci. 19, 1699 (2018).
Wittenbecher, C. et al. Plasma N-glycans as emerging biomarkers of cardiometabolic risk: a prospective investigation in the EPIC-Potsdam cohort study. Diabetes Care 43, 661–668 (2020).
Memarian, E. et al. Plasma protein N-glycosylation is associated with cardiovascular disease, nephropathy, and retinopathy in type 2 diabetes. BMJ Open. Diabetes Res. Care 9, e002345 (2021).
Testa, R. et al. N-glycomic changes in serum proteins in type 2 diabetes mellitus correlate with complications and with metabolic syndrome parameters. PLoS ONE 10, e0119983 (2015).
Gonzalez-Quesada, C. et al. Thrombospondin-1 induction in the diabetic myocardium stabilizes the cardiac matrix in addition to promoting vascular rarefaction through angiopoietin-2 upregulation. Circ. Res. 113, 1331–1344 (2013).
Chen, F. et al. Cardioprotective effect of decorin in type 2 diabetes. Front. Endocrinol. 11, 479258 (2020).
Lai, J. et al. Overexpression of decorin promoted angiogenesis in diabetic cardiomyopathy via IGF1R-AKT-VEGF signaling. Sci. Rep. 7, 44473 (2017).
Frangogiannis, N. G. Matricellular proteins in cardiac adaptation and disease. Physiol. Rev. 92, 635–688 (2012).
Way, K. J. et al. Expression of connective tissue growth factor is increased in injured myocardium associated with protein kinase C β2 activation and diabetes. Diabetes 51, 2709–2718 (2002).
Wang, X. et al. Adverse effects of high glucose and free fatty acid on cardiomyocytes are mediated by connective tissue growth factor. Am. J. Physiol. Cell Physiol. 297, C1490–C1500 (2009).
Twigg, S. M. Regulation and bioactivity of the CCN family of genes and proteins in obesity and diabetes. J. Cell Commun. Signal. 12, 359–368 (2018).
Accornero, F. et al. Genetic analysis of connective tissue growth factor as an effector of transforming growth factor β signaling and cardiac remodeling. Mol. Cell Biol. 35, 2154–2164 (2015).
Dorn, L. E., Petrosino, J. M., Wright, P. & Accornero, F. CTGF/CCN2 is an autocrine regulator of cardiac fibrosis. J. Mol. Cell. Cardiol. 121, 205–211 (2018).
Chatham, J. C., Young, M. E. & Zhang, J. Role of O-linked N-acetylglucosamine (O-GlcNAc) modification of proteins in diabetic cardiovascular complications. Curr. Opin. Pharmacol. 57, 1–12 (2021).
Hu, Y. et al. Adenovirus-mediated overexpression of O-GlcNAcase improves contractile function in the diabetic heart. Circ. Res. 96, 1006–1013 (2005).
Ramirez-Correa, G. et al. Removal of abnormal myofilament O-GlcNAcylation restores Ca2+ sensitivity in diabetic cardiac muscle. Diabetes 64, 3573–3587 (2015).
Hu, Y. et al. Increased enzymatic O-GlcNAcylation of mitochondrial proteins impairs mitochondrial function in cardiac myocytes exposed to high glucose. J. Biol. Chem. 284, 547–555 (2009).
Banerjee, P. S., Ma, J. & Hart, G. W. Diabetes-associated dysregulation of O-GlcNAcylation in rat cardiac mitochondria. Proc. Natl Acad. Sci. USA 112, 6050–6055 (2015).
Ma, J. et al. Comparative proteomics reveals dysregulated mitochondrial O-GlcNAcylation in diabetic hearts. J. Proteome Res. 15, 2254–2264 (2016).
Gawlowski, T. et al. Modulation of dynamin-related protein 1 (DRP1) function by increased O-linked-beta-N-acetylglucosamine modification (O-GlcNAc) in cardiac myocytes. J. Biol. Chem. 287, 30024–30034 (2012).
Cividini, F. et al. O-GlcNAcylation of 8-oxoguanine DNA glycosylase (Ogg1) impairs oxidative mitochondrial DNA lesion repair in diabetic hearts. J. Biol. Chem. 291, 26515–26528 (2016).
Hegyi, B., Bers, D. M. & Bossuyt, J. CaMKII signaling in heart diseases: emerging role in diabetic cardiomyopathy. J. Mol. Cell. Cardiol. 127, 246–259 (2019).
Erickson, J. R. et al. Diabetic hyperglycaemia activates CaMKII and arrhythmias by O-linked glycosylation. Nature 502, 372–376 (2013).
Lu, S. et al. Hyperglycemia acutely increases cytosolic reactive oxygen species via O-linked GlcNAcylation and CaMKII activation in mouse ventricular myocytes. Circ. Res. 126, e80–e96 (2020).
Kronlage, M. et al. O-GlcNAcylation of histone deacetylase 4 protects the diabetic heart from failure. Circulation 140, 580–594 (2019).
Xie, Z., He, C. & Zou, M. H. AMP-activated protein kinase modulates cardiac autophagy in diabetic cardiomyopathy. Autophagy 7, 1254–1255 (2011).
Sciarretta, S., Boppana, V. S., Umapathi, M., Frati, G. & Sadoshima, J. Boosting autophagy in the diabetic heart: a translational perspective. Cardiovasc. Diagn. Ther. 5, 394–402 (2015).
Wani, W. Y. et al. Regulation of autophagy by protein post-translational modification. Lab. Invest. 95, 14–25 (2015).
Zhang, J., Chatham, J. C. & Young, M. E. Circadian regulation of cardiac physiology: rhythms that keep the heart beating. Annu. Rev. Physiol. 82, 79–101 (2020).
Durgan, D. J. et al. O-GlcNAcylation, novel post-translational modification linking myocardial metabolism and cardiomyocyte circadian clock. J. Biol. Chem. 286, 44606–44619 (2011).
Qin, C. X. et al. Insights into the role of maladaptive hexosamine biosynthesis and O-GlcNAcylation in development of diabetic cardiac complications. Pharmacol. Res. 116, 45–56 (2017).
Aguilar, H. et al. Role for high-glucose-induced protein O-GlcNAcylation in stimulating cardiac fibroblast collagen synthesis. Am. J. Physiol. Cell Physiol. 306, C794–C804 (2014).
Ha, C. M. et al. Sustained increases in cardiomyocyte protein O-linked β-N-acetylglucosamine levels lead to cardiac hypertrophy and reduced mitochondrial function without systolic contractile impairment. J. Am. Heart Assoc. 12, e029898 (2023).
Snapp, K. R. et al. A novel P-selectin glycoprotein ligand-1 monoclonal antibody recognizes an epitope within the tyrosine sulfate motif of human PSGL-1 and blocks recognition of both P- and L-selectin. Blood 91, 154–164 (1998).
Hickey, M. J. et al. L-selectin facilitates emigration and extravascular locomotion of leukocytes during acute inflammatory responses in vivo. J. Immunol. 165, 7164–7170 (2000).
Rodgers, S. D., Camphausen, R. T. & Hammer, D. A. Sialyl Lewis(x)-mediated, PSGL-1-independent rolling adhesion on P-selectin. Biophys. J. 79, 694–706 (2000).
Sperandio, M. et al. α2,3-Sialyltransferase-IV is essential for L-selectin ligand function in inflammation. Eur. J. Immunol. 36, 3207–3215 (2006).
Yang, W. H., Nussbaum, C., Grewal, P. K., Marth, J. D. & Sperandio, M. Coordinated roles of ST3Gal-VI and ST3Gal-IV sialyltransferases in the synthesis of selectin ligands. Blood 120, 1015–1026 (2012).
Jung, U. & Ley, K. Mice lacking two or all three selectins demonstrate overlapping and distinct functions for each selectin. J. Immunol. 162, 6755–6762 (1999).
Nahrendorf, M. Myeloid cell contributions to cardiovascular health and disease. Nat. Med. 24, 711–720 (2018).
Randolph, G. J. The fate of monocytes in atherosclerosis. J. Thromb. Haemost. 7, 28–30 (2009).
Regal-McDonald, K. et al. Assessment of ICAM-1 N-glycoforms in mouse and human models of endothelial dysfunction. PLoS ONE 15, e0230358 (2020).
Regal-McDonald, K., Xu, B., Barnes, J. W. & Patel, R. P. High-mannose intercellular adhesion molecule-1 enhances CD16+ monocyte adhesion to the endothelium. Am. J. Physiol. Heart Circ. Physiol. 317, H1028–H1038 (2019).
Scott, D. W. et al. Role of endothelial N-glycan mannose residues in monocyte recruitment during atherogenesis. Arterioscler. Thromb. Vasc. Biol. 32, e51–e59 (2012).
Augustin-Voss, H. G. & Pauli, B. U. Migrating endothelial cells are distinctly hyperglycosylated and express specific migration-associated cell surface glycoproteins. J. Cell Biol. 119, 483–491 (1992).
Scott, D. W., Vallejo, M. O. & Patel, R. P. Heterogenic endothelial responses to inflammation: role for differentital N-glycosylation and vascular bed of origin. J. Am. Heart Assoc. 2, e000263 (2013).
Regal-McDonald, K. & Patel, R. P. Selective recruitment of monocyte subsets by endothelial N-glycans. Am. J. Pathol. 190, 947–957 (2020).
Aird, W. C. Phenotypic heterogeneity of the endothelium: II. Representative vascular beds. Circ. Res. 100, 174–190 (2007).
Scott, D. W., Vallejo, M. O. & Patel, R. P. Heterogenic endothelial responses to inflammation: role for differential N-glycosylation and vascular bed of origin. J. Am. Heart Assoc. 2, e000263 (2013).
Scott, D. W. & Patel, R. P. Endothelial heterogeneity and adhesion molecules N-glycosylation: implications in leukocyte trafficking in inflammation. Glycobiology 23, 622–633 (2013).
Sperandio, M., Gleissner, C. A. & Ley, K. Glycosylation in immune cell trafficking. Immunol. Rev. 230, 97–113 (2009).
Garcia-Vallejo, J. J. et al. Activation of human endothelial cells by tumor necrosis factor-α results in profound changes in the expression of glycosylation-related genes. J. Cell Physiol. 206, 203–210 (2006).
Chacko, B. K., Scott, D. W., Chandler, R. T. & Patel, R. P. Endothelial surface N-glycans mediate monocyte adhesion and are targets for anti-inflammatory effects of peroxisome proliferator-activated receptor γ ligands. J. Biol. Chem. 286, 38738–38747 (2011).
Mun, G. I., Jang, S. I. & Boo, Y. C. Laminar shear stress induces the expression of aquaporin 1 in endothelial cells involved in wound healing. Biochem. Biophys. Res. Commun. 430, 554–559 (2013).
Mun, G. I., Lee, S. J., An, S. M., Kim, I. K. & Boo, Y. C. Differential gene expression in young and senescent endothelial cells under static and laminar shear stress conditions. Free. Radic. Biol. Med. 47, 291–299 (2009).
Barallobre-Barreiro, J. et al. Extracellular matrix in vascular disease, part 2/4: JACC focus seminar. J. Am. Coll. Cardiol. 75, 2189–2203 (2020).
Ballout, R. A. & Remaley, A. T. GlycA: a new biomarker for systemic inflammation and cardiovascular disease (CVD) risk assessment. J. Lab. Precis. Med. 5, 17 (2020).
Akinkuolie, A. O., Buring, J. E., Ridker, P. M. & Mora, S. A novel protein glycan biomarker and future cardiovascular disease events. J. Am. Heart Assoc. 3, e001221 (2014).
Duprez, D. A. et al. Comparison of the predictive value of GlycA and other biomarkers of inflammation for total death, incident cardiovascular events, noncardiovascular and noncancer inflammatory-related events, and total cancer events. Clin. Chem. 62, 1020–1031 (2016).
Gruppen, E. G. et al. GlycA, a pro-inflammatory glycoprotein biomarker, and incident cardiovascular disease: relationship with C-reactive protein and renal function. PLoS ONE 10, e0139057 (2015).
Tian, C., Mahara, G., Zhang, H. & Tan, X. Association of immunoglobulin G N-glycosylation with carotid atherosclerotic plaque phenotypes and actual clinical cardiovascular events: a study protocol for a longitudinal prospective cohort study. BMJ Open. 12, e058922 (2022).
Radovani, B. et al. IgG N-glycosylation is altered in coronary artery disease. Biomolecules 13, 375 (2023).
Peng, J. et al. Supplementation with the sialic acid precursor N-acetyl-D-mannosamine breaks the link between obesity and hypertension. Circulation 140, 2005–2018 (2019).
Birukov, A. et al. Immunoglobulin G N-glycosylation signatures in incident type 2 diabetes and cardiovascular disease. Diabetes Care 45, 2729–2736 (2022).
Mise, K. et al. Novel urinary glycan biomarkers predict cardiovascular events in patients with type 2 diabetes: a multicenter prospective study with 5-year follow up (U-CARE Study 2). Front. Cardiovasc. Med. 8, 668059 (2021).
Li, X. et al. Type 2 diabetes mellitus is associated with the immunoglobulin G N-Glycome through putative proinflammatory mechanisms in an Australian population. OMICS 23, 631–639 (2019).
Wang, B. Y. et al. A nested case-control study to explore the association between immunoglobulin G N-glycans and ischemic stroke. Biomed. Env. Sci. 36, 389–396 (2023).
Plavsa, B. et al. The N-glycosylation of total plasma proteins and IgG in atrial fibrillation. Biomolecules 13, 605 (2023).
Romo, E. Z. & Zivkovic, A. M. Glycosylation of HDL-associated proteins and its implications in cardiovascular disease diagnosis, metabolism and function. Front. Cardiovasc. Med. 9, 928566 (2022).
Bolanle, I. O., Riches-Suman, K., Loubani, M., Williamson, R. & Palmer, T. M. Revascularisation of type 2 diabetics with coronary artery disease: insights and therapeutic targeting of O-GlcNAcylation. Nutr. Metab. Cardiovasc. Dis. 31, 1349–1356 (2021).
Byon, C. H. & Kim, S. W. Regulatory effects of O-GlcNAcylation in vascular smooth muscle cells on diabetic vasculopathy. J. Lipid Atheroscler. 9, 243–254 (2020).
Khanal, S. et al. Deletion of smooth muscle O-GlcNAc transferase prevents development of atherosclerosis in western diet-fed hyperglycemic ApoE-/- mice in vivo. Int. J. Mol. Sci. 24, 7899 (2023).
Xiong, X. et al. αSMA-Cre-mediated Ogt deletion leads to heart failure and vascular smooth muscle cell dysfunction in mice. Biochem. Biophys. Res. Commun. 625, 31–37 (2022).
Lima, V. V. et al. O-GlcNAcylation contributes to the vascular effects of ET-1 via activation of the RhoA/Rho-kinase pathway. Cardiovasc. Res. 89, 614–622 (2011).
Zhang, H. et al. Glutamine supplementation alleviated aortic atherosclerosis in mice model and in vitro. Proteomics 7, e2300179 (2023).
Hilgers, R. H. et al. Acute O-GlcNAcylation prevents inflammation-induced vascular dysfunction. Am. J. Physiol. Heart Circ. Physiol. 303, H513–H522 (2012).
Aulak, K. S. et al. Specific O-GlcNAc modification at Ser-615 modulates eNOS function. Redox Biol. 36, 101625 (2020).
Masaki, N. et al. O-GlcNAcylation mediates glucose-induced alterations in endothelial cell phenotype in human diabetes mellitus. J. Am. Heart Assoc. 9, e014046 (2020).
Zhang, L. et al. Oscillatory shear stress-mediated aberrant O-GlcNAc SIRT3 accelerates glycocalyx inflammatory injury via LKB1/p47(phox)/Hyal2 signaling. Cell Signal. 109, 110790 (2023).
Gorelik, A. et al. Genetic recoding to dissect the roles of site-specific protein O-GlcNAcylation. Nat. Struct. Mol. Biol. 26, 1071–1077 (2019).
Bennett, H. M., Stephenson, W., Rose, C. M. & Darmanis, S. Single-cell proteomics enabled by next-generation sequencing or mass spectrometry. Nat. Methods 20, 363–374 (2023).
Flynn, R. A. et al. Small RNAs are modified with N-glycans and displayed on the surface of living cells. Cell 184, 3109–3124.e22 (2021).
Yan, W. et al. Decorin gene delivery inhibits cardiac fibrosis in spontaneously hypertensive rats by modulation of transforming growth factor-β/Smad and p38 mitogen-activated protein kinase signaling pathways. Hum. Gene Ther. 20, 1190–1200 (2009).
Dang, H., Ye, Y., Zhao, X. & Zeng, Y. Identification of candidate genes in ischemic cardiomyopathy by gene expression omnibus database. BMC Cardiovasc. Disord. 20, 320 (2020).
Liu, C., Tong, H., Li, S. & Yan, Y. Effect of ECM2 expression on bovine skeletal muscle-derived satellite cell differentiation. Cell Biol. Int. 42, 525–532 (2018).
Andenaes, K. et al. The extracellular matrix proteoglycan fibromodulin is upregulated in clinical and experimental heart failure and affects cardiac remodeling. PLoS ONE 13, e0201422 (2018).
Hultgardh-Nilsson, A., Boren, J. & Chakravarti, S. The small leucine-rich repeat proteoglycans in tissue repair and atherosclerosis. J. Intern. Med. 278, 447–461 (2015).
Aboumsallem, J. P. et al. Multi-omics analyses identify molecular signatures with prognostic values in different heart failure aetiologies. J. Mol. Cell. Cardiol. 175, 13–28 (2023).
Zhu, Y., Yang, X. & Zu, Y. Integrated analysis of WGCNA and machine learning identified diagnostic biomarkers in dilated cardiomyopathy with heart failure. Front. Cell Dev. Biol. 10, 1089915 (2022).
Barallobre-Barreiro, J. et al. Proteomics analysis of cardiac extracellular matrix remodeling in a porcine model of ischemia/reperfusion injury. Circulation 125, 789–802 (2012).
Carlson, E. C. et al. Keratocan and lumican regulate neutrophil infiltration and corneal clarity in lipopolysaccharide-induced keratitis by direct interaction with CXCL1. J. Biol. Chem. 282, 35502–35509 (2007).
Malgija, B., Kumar, N. S. & Piramanayagam, S. Collective transcriptomic deregulation of hypertrophic and dilated cardiomyopathy – importance of fibrotic mechanism in heart failure. Comput. Biol. Chem. 73, 85–94 (2018).
Maiwald, S. et al. Mutation in KERA identified by linkage analysis and targeted resequencing in a pedigree with premature atherosclerosis. PLoS ONE 9, e98289 (2014).
Skenteris, N. T. et al. Osteomodulin attenuates smooth muscle cell osteogenic transition in vascular calcification. Clin. Transl. Med. 12, e682 (2022).
Goncalves, I. et al. Osteomodulin gene expression is associated with plaque calcification, stability, and fewer cardiovascular events in the CPIP cohort. Stroke 53, e79–e84 (2022).
Le Goff, M. M. et al. Opticin exerts its anti-angiogenic activity by regulating extracellular matrix adhesiveness. J. Biol. Chem. 287, 28027–28036 (2012).
Deckx, S. et al. Osteoglycin prevents the development of age-related diastolic dysfunction during pressure overload by reducing cardiac fibrosis and inflammation. Matrix Biol. 66, 110–124 (2018).
Petretto, E. et al. Integrated genomic approaches implicate osteoglycin (Ogn) in the regulation of left ventricular mass. Nat. Genet. 40, 546–552 (2008).
Fu, X. et al. Specialized fibroblast differentiated states underlie scar formation in the infarcted mouse heart. J. Clin. Invest. 128, 2127–2143 (2018).
Hessle, L. et al. The skeletal phenotype of chondroadherin deficient mice. PLoS ONE 8, e63080 (2014).
Hutter, R. et al. Novel small leucine-rich repeat protein podocan is a negative regulator of migration and proliferation of smooth muscle cells, modulates neointima formation, and is expressed in human atheroma. Circulation 128, 2351–2363 (2013).
Li, S. et al. Podocan promotes differentiation of bovine skeletal muscle satellite cells by regulating the Wnt4-β-catenin signaling pathway. Front. Physiol. 10, 1010 (2019).
Mochida, Y. et al. Podocan-like protein: a novel small leucine-rich repeat matrix protein in bone. Biochem. Biophys. Res. Commun. 410, 333–338 (2011).
Zhang, Z. et al. Whole-genome sequencing identifies novel candidate pathogenic variants associated with left ventricular non-compaction in a three-generation family. Clin. Transl. Med. 11, e501 (2021).
Koch, C. D., Lee, C. M. & Apte, S. S. Aggrecan in cardiovascular development and disease. J. Histochem. Cytochem. 68, 777–795 (2020).
Rauch, U., Feng, K. & Zhou, X. H. Neurocan: a brain chondroitin sulfate proteoglycan. Cell Mol. Life Sci. 58, 1842–1856 (2001).
Mishima, N. & Hoffman, S. Neurocan in the embryonic avian heart and vasculature. Anat. Rec. A Discov. Mol. Cell Evol. Biol. 272, 556–562 (2003).
Chelyshev, Y. A., Kabdesh, I. M. & Mukhamedshina, Y. O. Extracellular matrix in neural plasticity and regeneration. Cell Mol. Neurobiol. 42, 647–664 (2022).
Chen, J. et al. Endocan: a key player of cardiovascular disease. Front. Cardiovasc. Med. 8, 798699 (2021).
Klisic, A. & Patoulias, D. The role of endocan in cardiometabolic disorders. Metabolites 13, 640 (2023).
Solakyildirim, K. et al. Proteoglycan 4 (lubricin) is a highly sialylated glycoprotein associated with cardiac valve damage in animal models of infective endocarditis. Glycobiology 31, 1582–1595 (2021).
Das, N., Schmidt, T. A., Krawetz, R. J. & Dufour, A. Proteoglycan 4: from mere lubricant to regulator of tissue homeostasis and inflammation: does proteoglycan 4 have the ability to buffer the inflammatory response? Bioessays 41, e1800166 (2019).
Park, D. S. J. et al. Human pericardial proteoglycan 4 (lubricin): implications for postcardiotomy intrathoracic adhesion formation. J. Thorac. Cardiovasc. Surg. 156, 1598–1608 (2018).
Nahon, J. E. et al. Proteoglycan 4 regulates macrophage function without altering atherosclerotic lesion formation in a murine bone marrow-specific deletion model. Atherosclerosis 274, 120–127 (2018).
Huang, Q. & Huang, Q. Inhibition of lncRNA DANCR prevents heart failure by ameliorating cardiac hypertrophy and fibrosis via regulation of the miR-758-3p/PRG4/Smad axis. J. Cardiovasc. Transl. Res. 16, 1357–1372 (2023).
Lord, M. S., Melrose, J., Day, A. J. & Whitelock, J. M. The inter-α-trypsin inhibitor family: versatile molecules in biology and pathology. J. Histochem. Cytochem. 68, 907–927 (2020).
Edgell, C. J., BaSalamah, M. A. & Marr, H. S. Testican-1: a differentially expressed proteoglycan with protease inhibiting activities. Int. Rev. Cytol. 236, 101–122 (2004).
Nolan, D. K. et al. Fine mapping of a linkage peak with integration of lipid traits identifies novel coronary artery disease genes on chromosome 5. BMC Genet. 13, 12 (2012).
Choi, Y., Chung, H., Jung, H., Couchman, J. R. & Oh, E. S. Syndecans as cell surface receptors: unique structure equates with functional diversity. Matrix Biol. 30, 93–99 (2011).
Wang, X., Lu, Y., Xie, Y., Shen, J. & Xiang, M. Emerging roles of proteoglycans in cardiac remodeling. Int. J. Cardiol. 278, 192–198 (2019).
Sinha, A. et al. Protein-protein interactions between tenascin-R and RPTPζ/phosphacan are critical to maintain the architecture of perineuronal nets. J. Biol. Chem. 299, 104952 (2023).
Katraki-Pavlou, S. et al. Protein tyrosine phosphatase receptor-ζ1 deletion triggers defective heart morphogenesis in mice and zebrafish. Am. J. Physiol. Heart circ. Physiol. 322, H8–H24 (2022).
Zanin, M. K. et al. Distinct spatial and temporal distributions of aggrecan and versican in the embryonic chick heart. Anat. Rec. 256, 366–380 (1999).
Tamburini, E. et al. Structural deciphering of the NG2/CSPG4 proteoglycan multifunctionality. FASEB J. 33, 3112–3128 (2019).
Grako, K. A., Ochiya, T., Barritt, D., Nishiyama, A. & Stallcup, W. B. PDGF α-receptor is unresponsive to PDGF-AA in aortic smooth muscle cells from the NG2 knockout mouse. J. Cell Sci. 112, 905–915 (1999).
Alex, L., Tuleta, I., Harikrishnan, V. & Frangogiannis, N. G. Validation of specific and reliable genetic tools to identify, label, and target cardiac pericytes in mice. J. Am. Heart Assoc. 11, e023171 (2022).
Karmouch, J. et al. Distinct cellular basis for early cardiac arrhythmias, the cardinal manifestation of arrhythmogenic cardiomyopathy, and the skin phenotype of cardiocutaneous syndromes. Circ. Res. 121, 1346–1359 (2017).
Quijada, P. et al. Cardiac pericytes mediate the remodeling response to myocardial infarction. J. Clin. Invest. 133, e162188 (2023).
Strate, I., Tessadori, F. & Bakkers, J. Glypican4 promotes cardiac specification and differentiation by attenuating canonical Wnt and Bmp signaling. Development 142, 1767–1776 (2015).
Thota, L. N. R. & Chignalia, A. Z. The role of the glypican and syndecan families of heparan sulfate proteoglycans in cardiovascular function and disease. Am. J. Physiol. Cell Physiol. 323, C1052–C1060 (2022).
Melleby, A. O. et al. The heparan sulfate proteoglycan glypican-6 is upregulated in the failing heart, and regulates cardiomyocyte growth through ERK1/2 signaling. PLoS ONE 11, e0165079 (2016).
Souza, D. S., Chignalia, A. Z. & Carvalho-de-Souza, J. L. Modulation of cardiac voltage-activated K+ currents by glypican 1 heparan sulfate proteoglycan. Life Sci. 308, 120916 (2022).
Nonaka, R. et al. Perlecan deficiency causes endothelial dysfunction by reducing the expression of endothelial nitric oxide synthase. Physiol. Rep. 3, e12272 (2015).
Sasse, P. et al. Perlecan is critical for heart stability. Cardiovasc. Res. 80, 435–444 (2008).
Bassat, E. et al. The extracellular matrix protein agrin promotes heart regeneration in mice. Nature 547, 179–184 (2017).
Sun, X. et al. The extracellular matrix protein agrin is essential for epicardial epithelial-to-mesenchymal transition during heart development. Development 148, dev197525 (2021).
Baehr, A. et al. Agrin promotes coordinated therapeutic processes leading to improved cardiac repair in pigs. Circulation 142, 868–881 (2020).
Isobe, K. et al. Inhibition of endostatin/collagen XVIII deteriorates left ventricular remodeling and heart failure in rat myocardial infarction model. Circ. J. 74, 109–119 (2010).
Moulton, K. S. et al. Loss of collagen XVIII enhances neovascularization and vascular permeability in atherosclerosis. Circulation 110, 1330–1336 (2004).
Rasi, K. et al. Collagen XV is necessary for modeling of the extracellular matrix and its deficiency predisposes to cardiomyopathy. Circ. Res. 107, 1241–1252 (2010).
Eklund, L. et al. Lack of type XV collagen causes a skeletal myopathy and cardiovascular defects in mice. Proc. Natl Acad. Sci. USA 98, 1194–1199 (2001).
Durgin, B. G. et al. Smooth muscle cell-specific deletion of Col15a1 unexpectedly leads to impaired development of advanced atherosclerotic lesions. Am. J. Physiol. Heart Circ. Physiol. 312, H943–H958 (2017).
van der Rest, M. & Mayne, R. Type IX collagen proteoglycan from cartilage is covalently cross-linked to type II collagen. J. Biol. Chem. 263, 1615–1618 (1988).
Izu, Y. & Birk, D. E. Collagen XII mediated cellular and extracellular mechanisms in development, regeneration, and disease. Front. Cell Dev. Biol. 11, 1129000 (2023).
Liu, C. Y., Olsen, B. R. & Kao, W. W. Developmental patterns of two α1(IX) collagen mRNA isoforms in mouse. Dev. Dyn. 198, 150–157 (1993).
Marro, J., Pfefferli, C., de Preux Charles, A. S., Bise, T. & Jazwinska, A. Collagen XII contributes to epicardial and connective tissues in the zebrafish heart during ontogenesis and regeneration. PLoS ONE 11, e0165497 (2016).
Gil-Cayuela, C. et al. New altered non-fibrillar collagens in human dilated cardiomyopathy: role in the remodeling process. PLoS ONE 11, e0168130 (2016).
Peacock, J. D., Lu, Y., Koch, M., Kadler, K. E. & Lincoln, J. Temporal and spatial expression of collagens during murine atrioventricular heart valve development and maintenance. Dev. Dyn. 237, 3051–3058 (2008).
Williams, B. et al. Use of whole genome analysis to identify shared genomic variants across breeds in canine mitral valve disease. Hum. Genet. 140, 1563–1568 (2021).
Isono, T. et al. Transcriptome analysis of a dog model of congestive heart failure shows that collagen-related 2-oxoglutarate-dependent dioxygenases contribute to heart failure. Sci. Rep. 12, 22569 (2022).
Imoto-Tsubakimoto, H. et al. Serglycin is a novel adipocytokine highly expressed in epicardial adipose tissue. Biochem. Biophys. Res. Commun. 432, 105–110 (2013).
Ilgin, B. U. et al. Association between serum serglycin levels and ST-segment elevation myocardial infarction. Arq. Bras. Cardiol. 116, 756–762 (2021).
Zhuo, L., Salustri, A. & Kimata, K. A physiological function of serum proteoglycan bikunin: the chondroitin sulfate moiety plays a central role. Glycoconj. J. 19, 241–247 (2002).
Maurel, P., Rauch, U., Flad, M., Margolis, R. K. & Margolis, R. U. Phosphacan, a chondroitin sulfate proteoglycan of brain that interacts with neurons and neural cell-adhesion molecules, is an extracellular variant of a receptor-type protein tyrosine phosphatase. Proc. Natl Acad. Sci. USA 91, 2512–2516 (1994).
Wen, W., Moses, M. A., Wiederschain, D., Arbiser, J. L. & Folkman, J. The generation of endostatin is mediated by elastase. Cancer Res. 59, 6052–6056 (1999).
Wen, J. et al. Xylose phosphorylation functions as a molecular switch to regulate proteoglycan biosynthesis. Proc. Natl Acad. Sci. USA 111, 15723–15728 (2014).
Acknowledgements
This work was supported in part by the National Institutes of Health (NIH) R21HL152354 and R01HL149354 to J.C.C. and HL153113 to R.P.P. The authors are grateful for the many discussions with A. R. Wende (University of Alabama at Birmingham, AL, USA) during the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Both authors contributed substantially to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Cardiology thanks Ivan Gudelj, Gordon Lauc and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chatham, J.C., Patel, R.P. Protein glycosylation in cardiovascular health and disease. Nat Rev Cardiol (2024). https://doi.org/10.1038/s41569-024-00998-z
Accepted:
Published:
DOI: https://doi.org/10.1038/s41569-024-00998-z